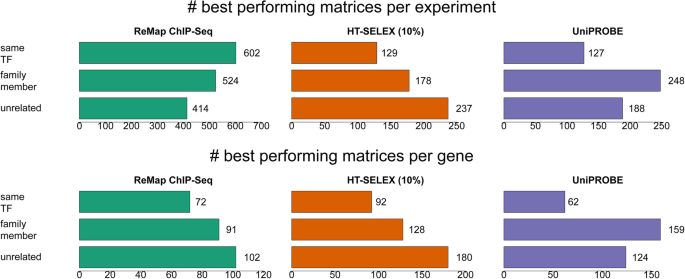
Insights gained from a comprehensive all-against-all transcription factor binding motif benchmarking study | Genome Biology | Full Text
![PDF] Cistrome Data Browser: a data portal for ChIP-Seq and chromatin accessibility data in human and mouse | Semantic Scholar PDF] Cistrome Data Browser: a data portal for ChIP-Seq and chromatin accessibility data in human and mouse | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/68bd3de081d1baa8df24bec6ec2ad46df5e48b6c/5-Figure2-1.png)
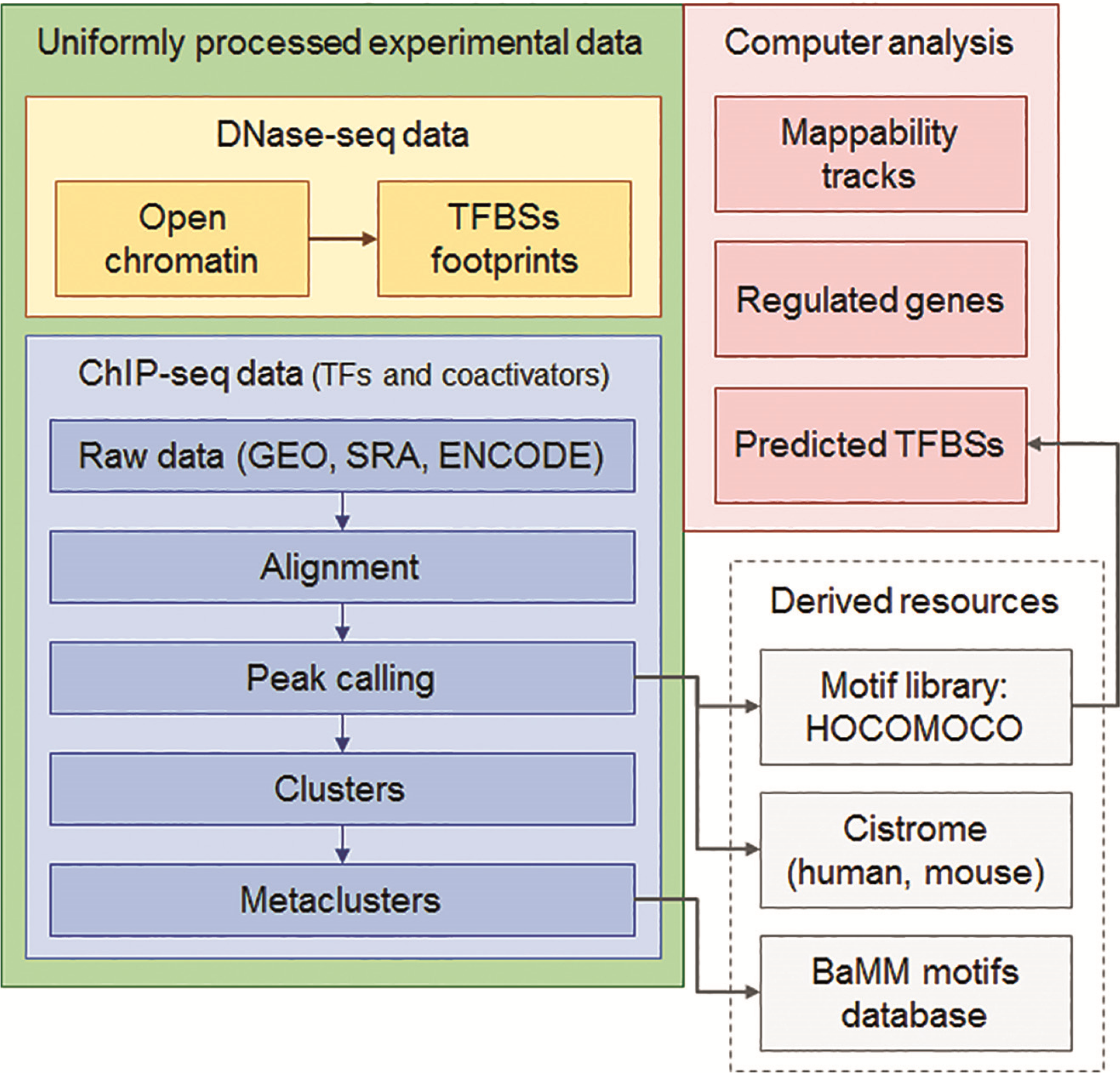
PDF] Cistrome Data Browser: a data portal for ChIP-Seq and chromatin accessibility data in human and mouse | Semantic Scholar

Quantification of Differential Transcription Factor Activity and Multiomics-Based Classification into Activators and Repressors: diffTF - ScienceDirect
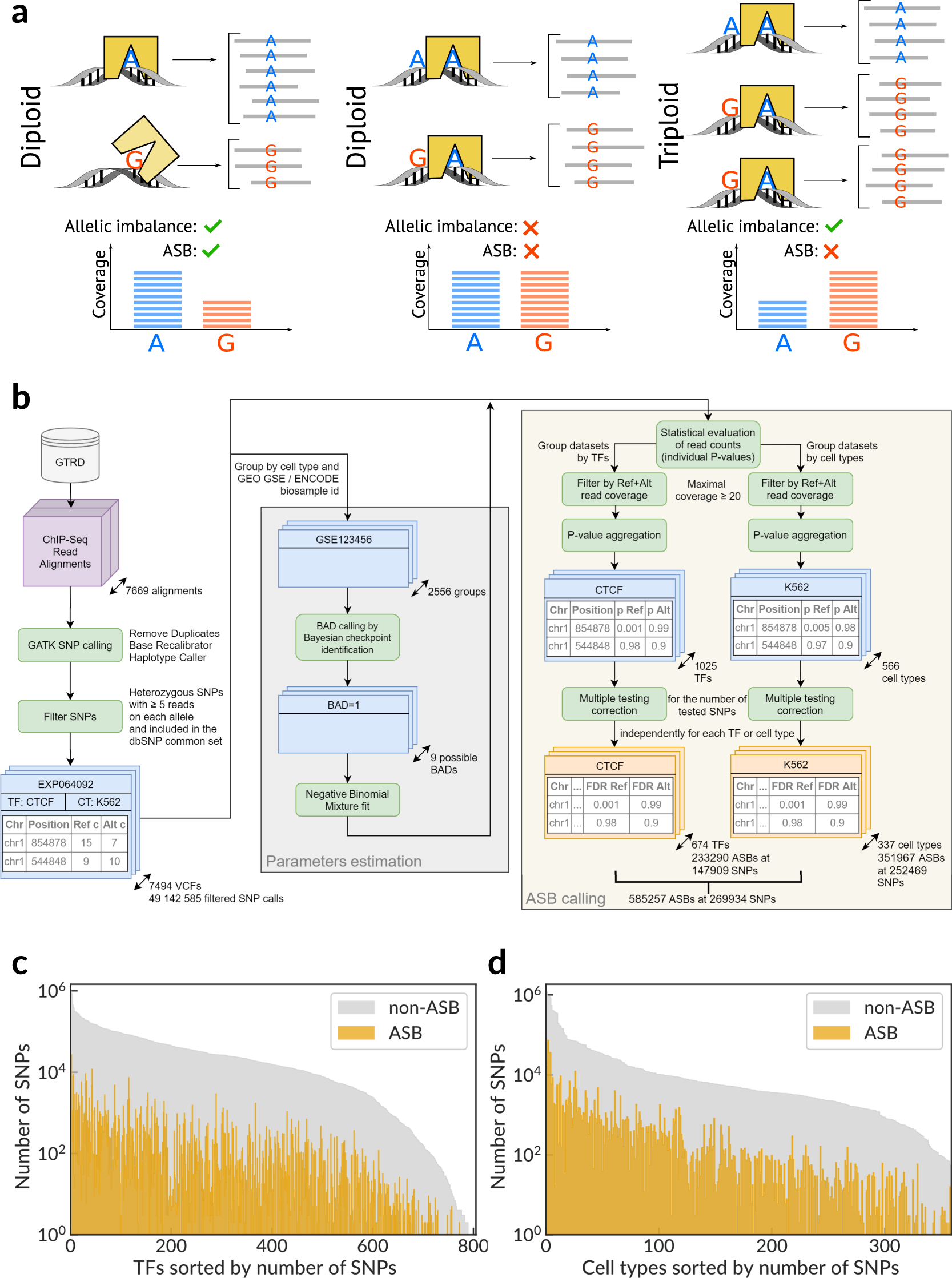
Landscape of allele-specific transcription factor binding in the human genome | Nature Communications

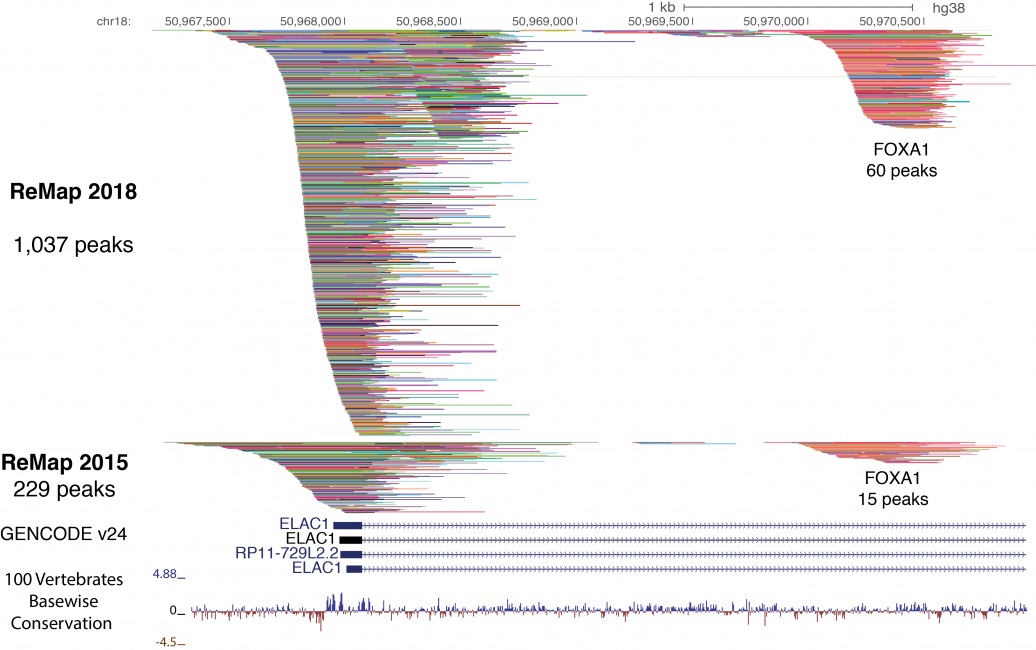
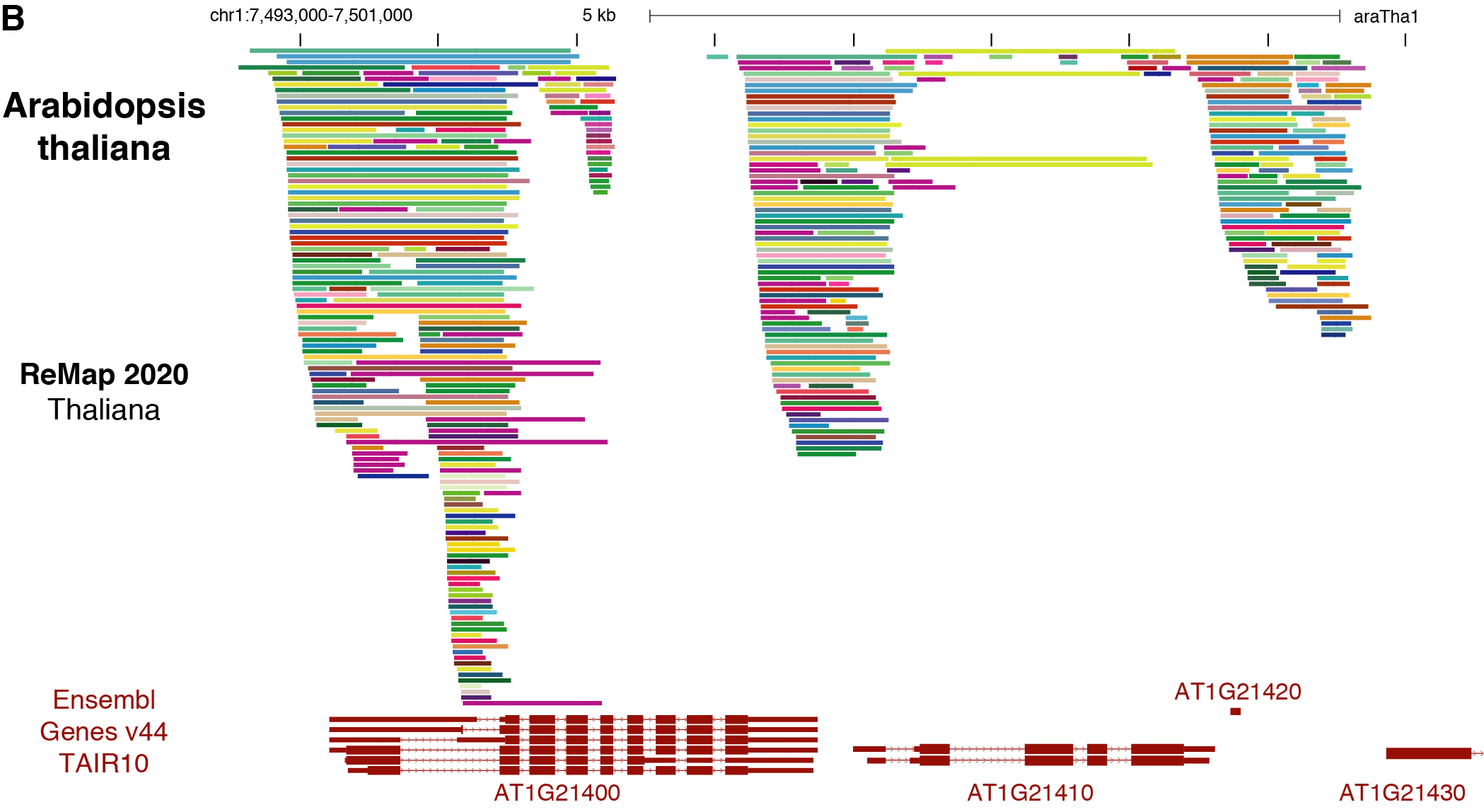
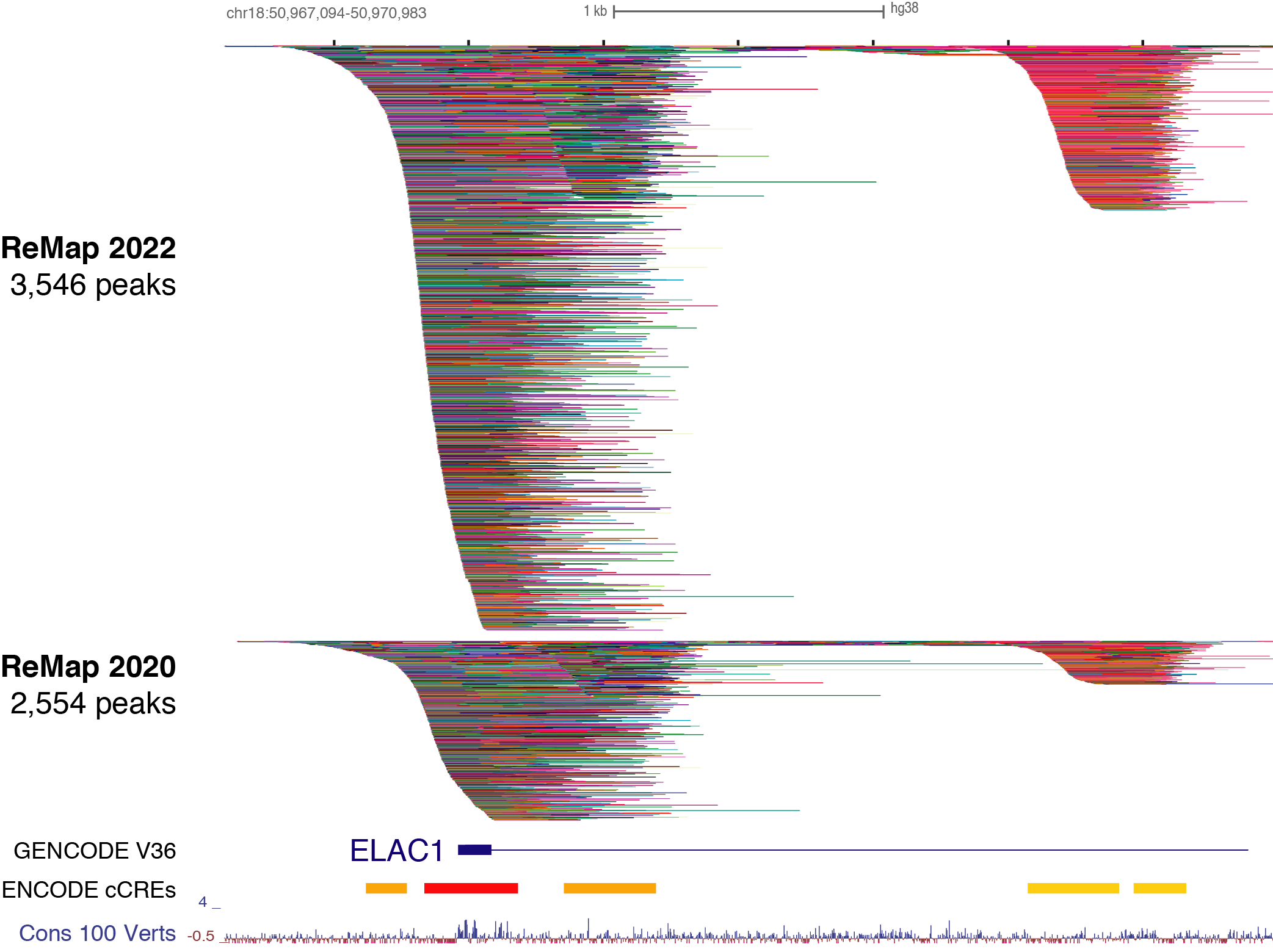
ReMap 2020: a database of regulatory regions from an integrative analysis of Human and Arabidopsis DNA-binding sequencing experiments. - Abstract - Europe PMC

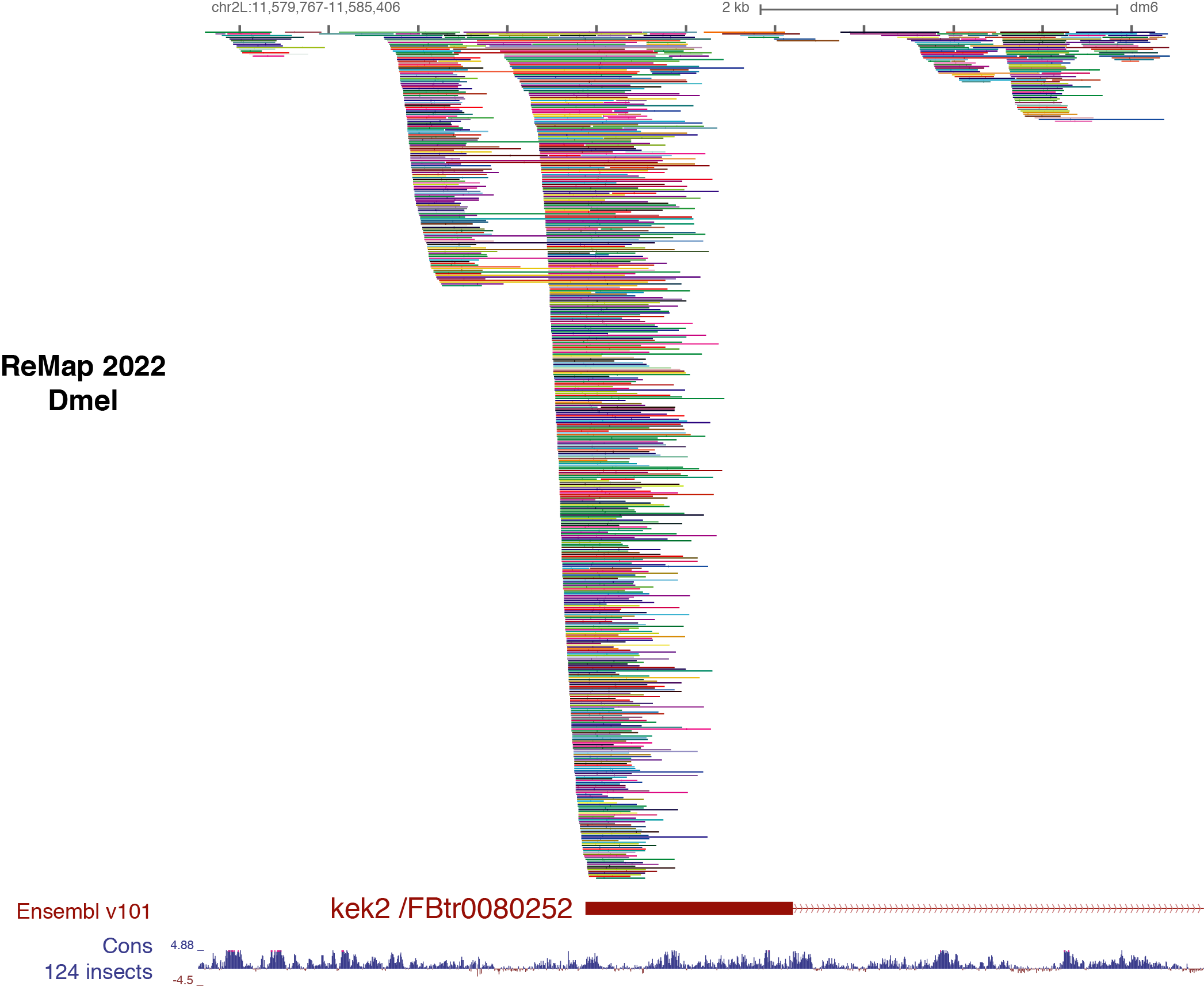
Benoit Ballester on Twitter: "The ReMap 2022 release is now on-line at https://t.co/3bv9RdYRfn. Regulatory ChIP-seq catalogue have been updated for Human and Arabidopsis, with new Mouse and Drosophila regulatory catalogues. https://t.co/m6jwUPyucS" /

Overview of MANTA2. a) Intersection of the ReMap ChIP-seq regions with... | Download Scientific Diagram
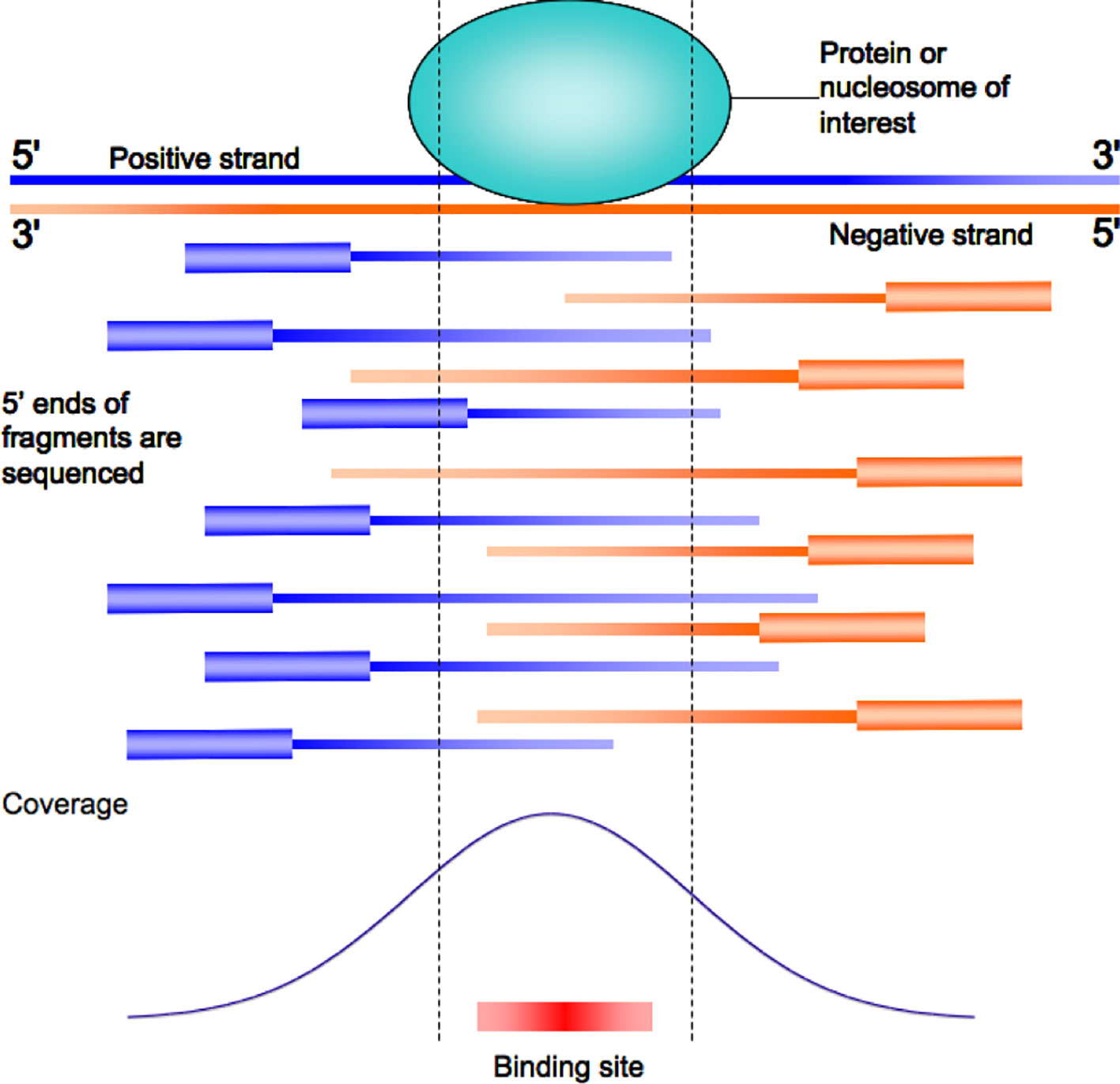
Frontiers | Integrating Peak Colocalization and Motif Enrichment Analysis for the Discovery of Genome-Wide Regulatory Modules and Transcription Factor Recruitment Rules
![Predicting stimulation-dependent enhancer-promoter interactions from ChIP- Seq time course data [PeerJ] Predicting stimulation-dependent enhancer-promoter interactions from ChIP- Seq time course data [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2017/3742/1/fig-1-full.png)
Predicting stimulation-dependent enhancer-promoter interactions from ChIP- Seq time course data [PeerJ]
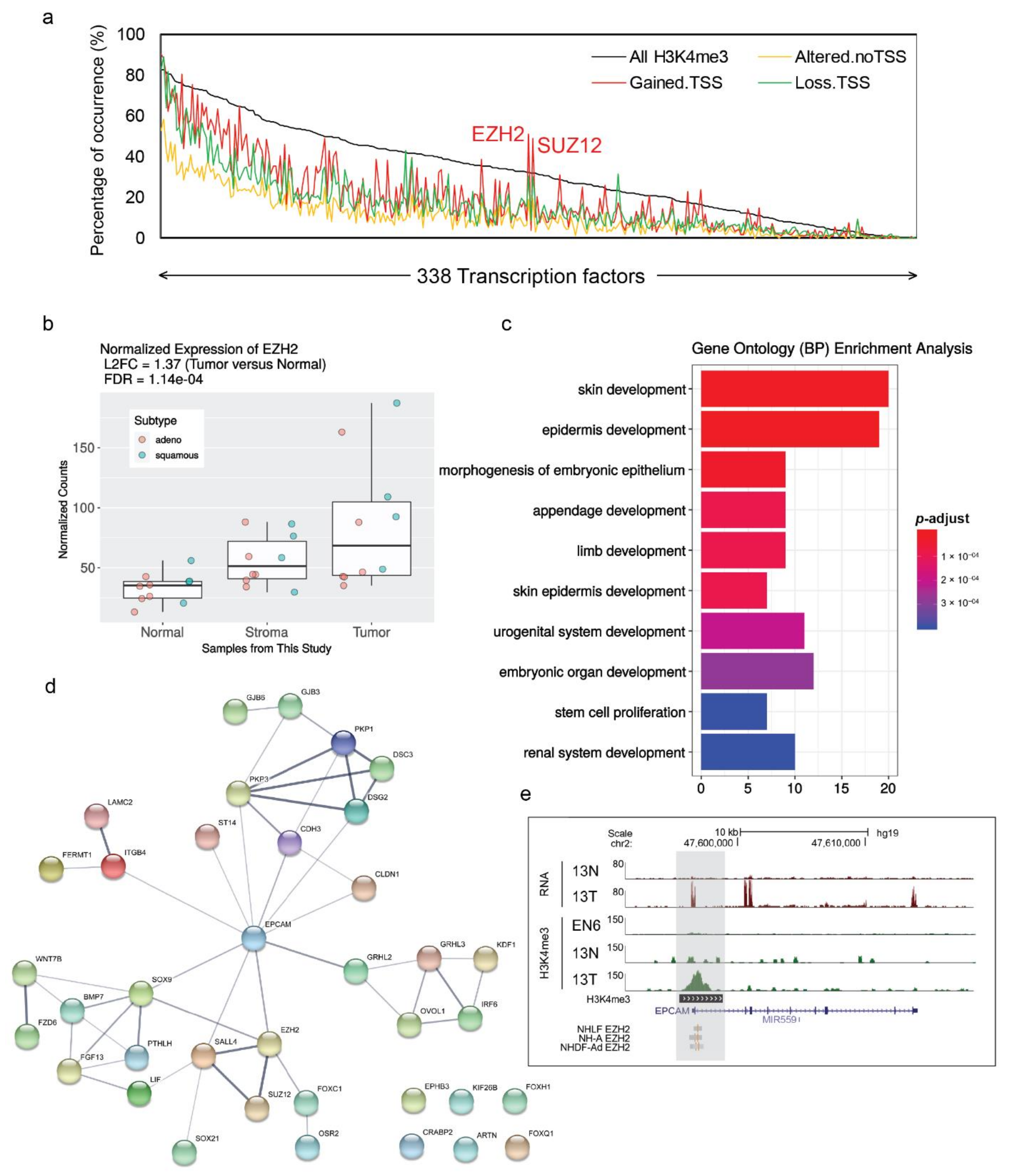
Cancers | Free Full-Text | Integrative RNA-Seq and H3 Trimethylation ChIP- Seq Analysis of Human Lung Cancer Cells Isolated by Laser-Microdissection